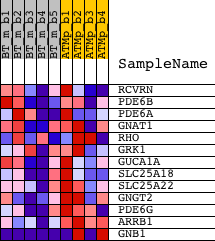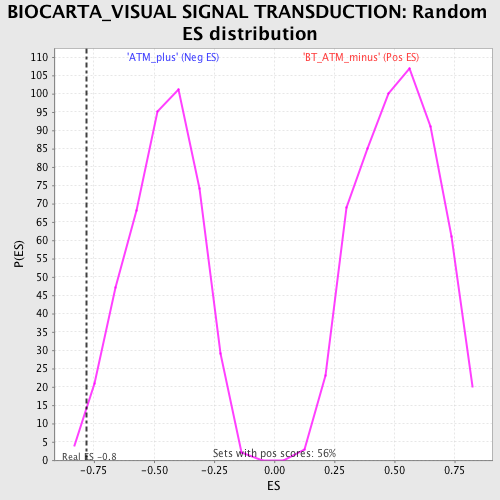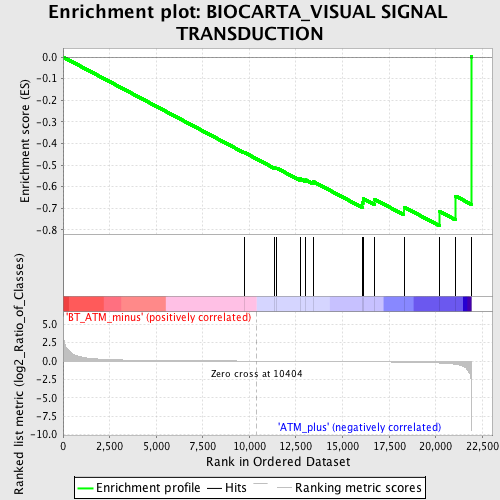
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_02_BT_ATM_minus_versus_ATM_plus.phenotype_BT_ATM_minus_versus_ATM_plus.cls #BT_ATM_minus_versus_ATM_plus.phenotype_BT_ATM_minus_versus_ATM_plus.cls #BT_ATM_minus_versus_ATM_plus_repos |
| Phenotype | phenotype_BT_ATM_minus_versus_ATM_plus.cls#BT_ATM_minus_versus_ATM_plus_repos |
| Upregulated in class | ATM_plus |
| GeneSet | BIOCARTA_VISUAL SIGNAL TRANSDUCTION |
| Enrichment Score (ES) | -0.7819688 |
| Normalized Enrichment Score (NES) | -1.6899754 |
| Nominal p-value | 0.013605442 |
| FDR q-value | 0.2776591 |
| FWER p-Value | 0.97 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | RCVRN | 1450215_at | 9736 | 0.009 | -0.4419 | No | ||
| 2 | PDE6B | 1419740_at | 11366 | -0.013 | -0.5130 | No | ||
| 3 | PDE6A | 1450415_at | 11482 | -0.014 | -0.5146 | No | ||
| 4 | GNAT1 | 1460212_at | 12723 | -0.030 | -0.5634 | No | ||
| 5 | RHO | 1425171_at 1425172_at 1451617_at 1451618_at | 13038 | -0.035 | -0.5689 | No | ||
| 6 | GRK1 | 1421361_at | 13436 | -0.041 | -0.5767 | No | ||
| 7 | GUCA1A | 1421061_at | 16063 | -0.084 | -0.6751 | No | ||
| 8 | SLC25A18 | 1428964_at | 16107 | -0.084 | -0.6555 | No | ||
| 9 | SLC25A22 | 1452653_at | 16730 | -0.098 | -0.6588 | No | ||
| 10 | GNGT2 | 1428733_at | 18305 | -0.142 | -0.6943 | No | ||
| 11 | PDE6G | 1425100_a_at 1450453_a_at | 20228 | -0.261 | -0.7152 | Yes | ||
| 12 | ARRB1 | 1460444_at | 21076 | -0.432 | -0.6435 | Yes | ||
| 13 | GNB1 | 1417432_a_at 1425908_at 1454696_at 1459442_at | 21915 | -2.671 | 0.0009 | Yes |